Negative stain- and cryo-electron microscopy
TEM Glacios 200kV: Falcon 4i Camera + Selectris X Energy Filter
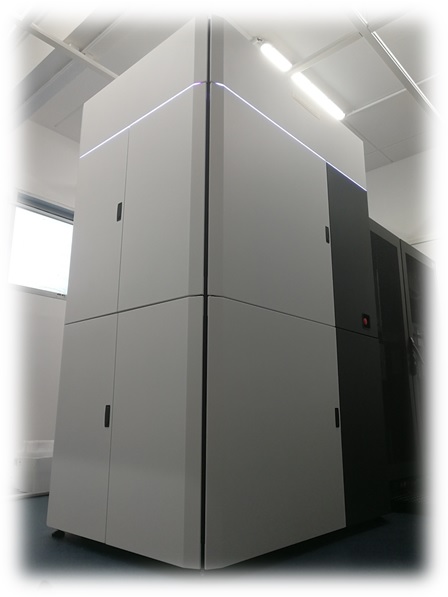

TEM Glacios at the AFMB lab
Cryo-EM field has been developing with unprecedented pace. Modern TEM allows collect data with increasing speed and high efficient robotic systems integrated in the microscope enable multiple experiments that can be done in fully automated non-assisted fashion. The EM workflow on modern microscopes is perfectly reliable and optimally set for quick evaluation of sample quality and setting up data acquisition.
We welcome users from any life sciences labs. Assistance is offered during sample preparation and imaging sessions. Users can benefit from a prep lab at AFMB if any additional preparations need to be done to the samples. For grid freezing there are Vitrobot Mark4 and Leica GP available to users.
Advanced EM users can also benefit from training offered by the platform. The training program can be tailored to specific user demands.
While high-end microscope is available at our facility we strongly recommend users to test their samples by negative staining EM on a Tecnai 120 that is also available at our lab.
No prior assessment of projects is necessary to access our facility but we do advise to get in touch with the facility manager to discuss the project before planning cryo-EM work.
Publications citing this service
- Nilsson J, Baroudi H, Gondelaud F, Pesce G, Bignon C, Ptchelkine D, Chamiech J, Cottet Hervé, Kajava A, Longhi S (2023) Molecular Determinants of Fibrillation in a Viral Amyloidogenic Domain from Combined Biochemical and Biophysical Studies. Int J Mol Sci 24:399
- Pesce G, Gondelaud F, Ptchelkine D, Nilsson J, Bignon C, Cartalas J, Fourquet P, Longhi S (2022) Experimental Evidence of Intrinsic Disorder and Amyloid Formation by the Henipavirus W Proteins Int J Mol Sci 23:923
- Gondelaud F, Pesce G, Nilsson J, Bignon C, Ptchelkine D, Gerlier D, Mathieu C, Longhi S (2022) Functional benefit of structural disorder for the replication of measles, Nipah and Hendra viruses. Essays in Bioc 66:915-934
- Bebeacua C, Tremblay D, Farenc C, Chapot-Chartier MP, Sadovskaya I, van Heel M, Veesler D, Moineau S, Cambillau C (2013) Structure, adsorption to host, and infection mechanism of virulent lactococcal phage p2. J Virol 87:12302-12.
- Desmyter A, Farenc C, Mahony J, Spinelli S, Bebeacua C, Blangy S, Veesler D, van Sinderen D, Cambillau C (2013) Viral infection modulation and neutralization by camelid nanobodies. Proc Natl Acad Sci U S A 110:E1371-9.
- Le Breton M, Meyniel-Schicklin L, Deloire A, Coutard B, Canard B, de Lamballerie X, Andre P, Rabourdin-Combe C, Lotteau V, Davoust N (2011) Flavivirus NS3 and NS5 proteins interaction network : a high-throughput yeast two-hybrid screen. BMC Microbiol 11, 234.
- Sciara G, Bebeacua C, Bron P, Tremblay D, Ortiz-Lombardia M, Lichière J, van Heel M, Campanacci V, Moineau S, Cambillau C (2010) Structure of lactococcal phage p2 baseplate and its mechanism of activation. Proc Natl Acad Sci USA 10:6852-7.





